| File Compression and Expansion of the Genetic Code by the Use of the
Yin/Yang Directions to Find its Sphered Cube |
| Fernando Castro Chavez* |
| Department of Medicine, Atherosclerosis and Vascular Medicine Section, Baylor College of Medicine, Houston, TX, USA |
| Corresponding Author : |
Fernando Castro Chavez
Department of Medicine
Atherosclerosis and Vascular Medicine Section
Baylor College of Medicine, Houston, TX, USA
Tel: +1 713 798 4177
Fax: 713-798-4121
E-mail: fdocc@yahoo.com |
| |
| Received March 04, 2014; Accepted May 20, 2014; Published June 05, 2014 |
| |
|
Citation: Chavez FC (2014) File Compression and Expansion of the Genetic Code by the Use of the Yin/Yang Directions to Find its Sphered Cube. J Biodivers Biopros Dev 1:112. doi: 10.4172/ijbbd.1000112 |
| |
|
Copyright: © 2014 Chavez FC. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. |
| |
Related article at
 Pubmed Pubmed  Scholar Google Scholar Google |
| |
Visit for more related articles at
 Journal of Biodiversity, Bioprospecting and Development Journal of Biodiversity, Bioprospecting and Development |
| |
| Abstract |
| |
Objective: The objective of this article is to demonstrate that the genetic code can be studied and represented in a 3-D Sphered Cube for bioinformatics and for education by using the graphical help of the ancient “Book of Changes” or I Ching for the comparison, pair by pair, of the three basic characteristics of nucleotides: H-bonds, molecular structure, and their tautomerism. Methods: The source of natural biodiversity is the high plasticity of the genetic code, analyzable with a reverse engineering of its 2-D and 3-D representations (here illustrated), but also through the classical 64-hexagrams of the ancient I Ching, as if they were the 64-codons or words of the genetic code. Results: In this article, the four elements of the Yin/Yang were found by correlating the 3x2=6 sets of Cartesian comparisons of the mentioned properties of nucleic acids, to the directionality of their resulting blocks of codons grouped according to their resulting amino acids and/or functions, integrating a 384 codon Sphered Cube whose function is illustrated by comparing six brain peptides and a promoter of osteoblasts from Humans versus Neanderthal, as well as to Negadi’s work on the importance of the number 384 within the genetic code. Conclusions: Starting with the codon/anticodon correlation of Nirenberg, published in full here for the first time, and by studying the genetic code and its 3-D display, the buffers of reiteration within codons codifying for the same amino acid, displayed the two long (binary number one) and older Yin/Yang arrows that travel in opposite directions, mimicking the parental DNA strands, while annealing to the two younger and broken (binary number zero) Yin/Yang arrows, mimicking the new DNA strands; the graphic analysis of the genetic code and its plasticity was helpful to compare compatible sequences, while further exploring the wondrous biodiversity of nature for educational purposes. |
| |
| Keywords |
| |
| Genetic code; I Ching; Cube; Sphered cube; Cubed
sphere; Yin and yang; Bioinformatics |
| |
| Introduction |
| |
| When studying the genetic code, the first question to ask could be:
“What is a ‘code’?” The most obvious answer is that a code is a set of
correspondences between two different languages. In the same way,
the bioinformatics context of this article is an interface between the
geometry within bioinformatics and its cryptography [1]. |
| |
| For example, let’s remember the intuitive letter-to-number code
of our childhood (evocative of the Roman numerals that lacked of
zero), also called the “letter number” cipher or code, in which the
letters of an alphabet are replaced by numbers in their respective order:
transforming abc… into 1-2-3…1 |
| |
| A more counter-intuitive variant of such letter-to-number code is
the one that includes the zero (0) at the start (as did the Arabic numerical
system, now in worldwide use, even in programming, a system that
apparently was developed independently by the Mayans); starting with
abc…, which in this new code that includes the zero, is transformed
into 0-1-2…, ending with …xyz being translated into …23-24-25 [1];
and, even the fingers of both of our hands can be used in mnemonics in order to remember this code, and by analogy, to remember the genetic
code; furthermore, our four extremities can be used to remember its
four foundational nucleotides (A, T, C, G)2 for a better understanding
of the bio-flexibility of the genetic code. |
| |
| Thus far, these are the basic aspects on the study of the genetic code,
aspects relatively easy to remember by a student exposed for the first
time to it; but we are going to find here additional tools that can be
used, not only in education, but also in bioinformatics, for the more
deeply appreciation of the source of information contained within the
genetic code. For the basic computational concept of file compression,
we read: “File compression reduces the size of a file by cleverly taking
out parts of the contents of the file that aren’t needed... Zipping is very
common, particularly because it reduces the amount of data that needs
to be transported from here to there” [2]. |
| |
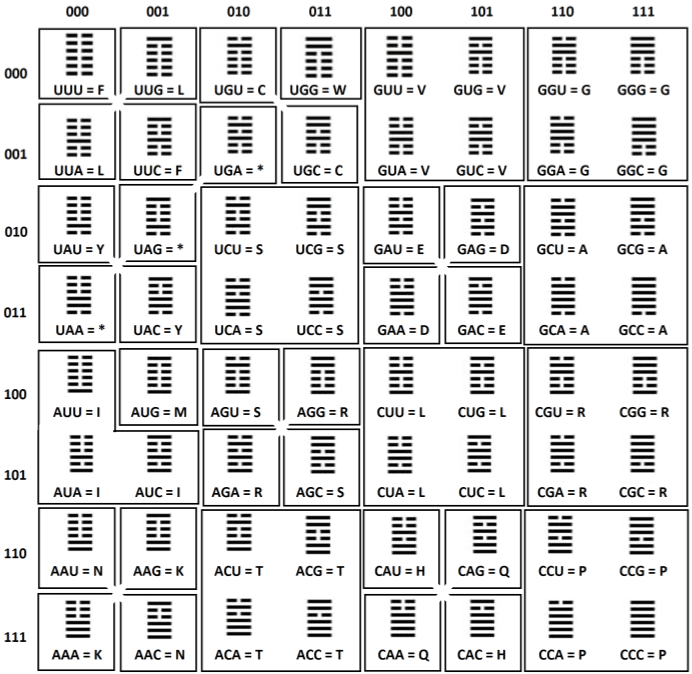
| We need to start with the unpublished and handwritten
representation of the set of 64 codons done by Nirenberg in 1965 [3]
(shown in Table 1; published in full as a primer in this article). After the representation of Nirenberg (Table 1), the table of Crick arrived in
1967 [4], then the circular representation by Bresch and Hausmann in
1972 [5], and then Fujimoto’s tetrahedron in 1987 [6], being the last
one similar to a backbone developed by Kepler many years before [7];
then, a set of more recent rules of variation [8,9] was discovered while
carefully analyzing [5], plus newer representations, including the use
of the I Ching [10], the tetrahedron [11], and the rotating square [12],
among others. |
| |
| Efforts have been done to represent the genetic code in a binary
way independent of the I Ching [13]; however, the I Ching is useful for
the study the genetic code and was previously compared to a detailed
treatment of Nirenberg’s Table in order to obtain quasi-symmetrical
representations of chromosomes [10]. Likewise, in a similar fashion
that the currently known letters of the genetic code are represented by
AGTC, the first Hebrew mention of the genetic code3 uses four letters
(grouped in 2×2), exactly as it happens in real life: being the four letters
of the genetic code grouped by threes to produce 4×4×4=64 codons or
words of exactly the same length per codon (three letters). |
| |
| Now, the purpose of a teacher of the genetic code is to make it known by illustrating its extremely intelligent design and its
complexities in the simplest way as possible; however, most of the
current text books spend only one page representing the genetic
code via Crick’s ‘square’ representation [4]. The word ‘genetic’
can be defined as: ‘the biological information that specifies the
characteristics and metabolism of an organism’. The information
in most of the biological life on earth is contained inside the nowfamous
double helix [14] that swirls like in a ‘fusion dance’, tightly
packaged within the chromosomes inside the nucleus of each living
cell, except for the red blood cells that, by design, expel their nucleus,
transforming themselves into biological ‘automatons’, or ‘robots’, for
the cellular exchange between oxygen and CO2. |
| |
| The genetic information that is transcribed from DNA to RNA in
the nucleus is translated into proteins in the cellular cytoplasm, ending
forming, these proteins, the structures and metabolism of our bodies. |
| |
| The basic units that contain the information for our proteins are
called ‘genes’, with each gene being composed of multiple nucleotides.
The modular structure of the genes integrated by exons depicts the
potential of a gene to produce several proteins through the ‘cut-andpaste’
(splicing) mechanism. |
| |
| The words of the genetic code are the groups of three nucleotides
called codons, producing the proteins that are formed by different
combinations of the 20 amino acids, with some of those combinations
being so frequent that these popular groups of amino acids, or of
peptides, are considered as ‘domains’. |
| |
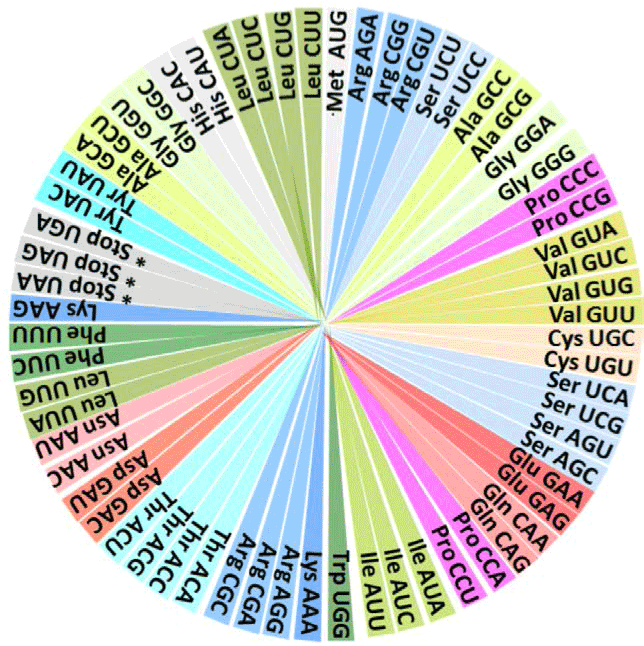
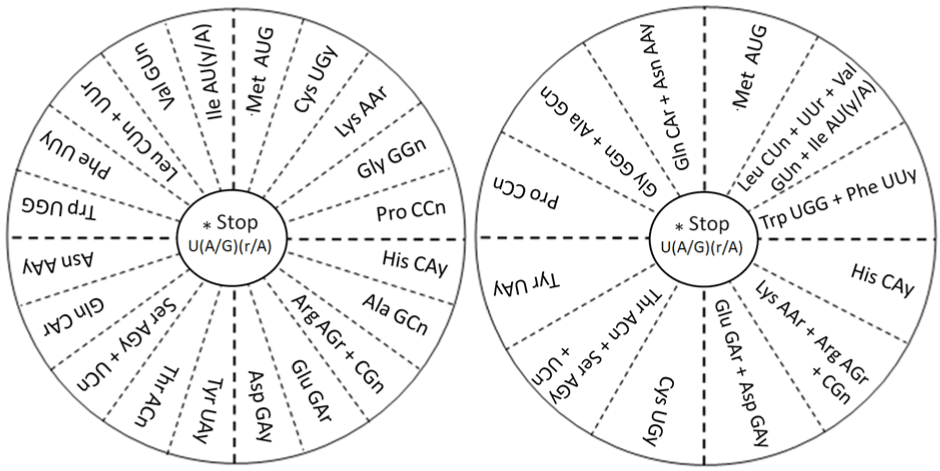
| Next, the reverse-engineered Figures 1 and 2 are the square and
the circular representations, respectively, of a functional tetrahedron
discovered earlier [11] as a representation of the genetic code. |
| |
| The periodicities or rotational correspondences of these
representations of the genetic code are clearly identifiable when we
rotate the quadrants by 90 degrees, keeping a positional and prominent
correspondence for most of the essential hydrophobic amino acids
located at the centre of the square of Figure 1 [12], remaining at the same relative rotational position in the circle shown in Figure 2 [8,9]. |
| |
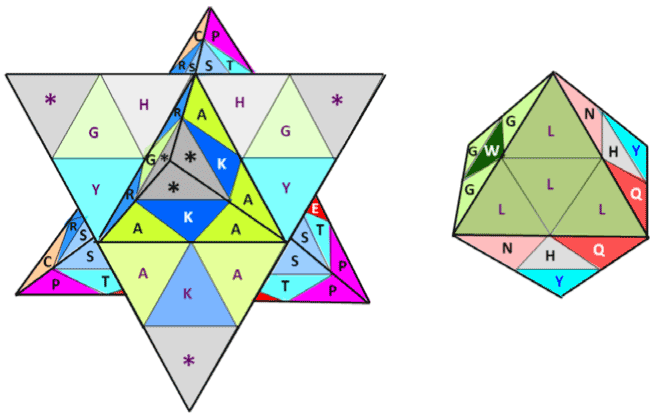
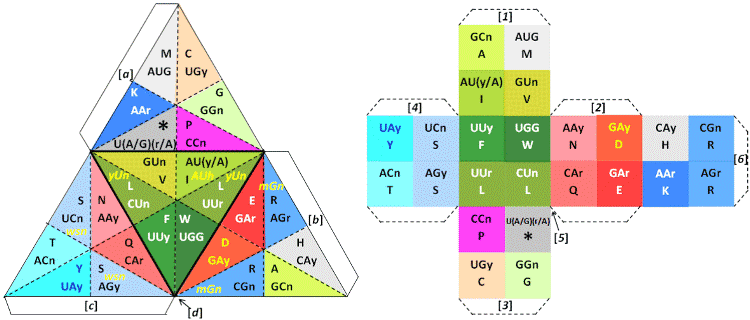
| Also, and in order to detail the representation of the genetic code
introduced in [10], the folded and ‘compressed’ 3-D representation of a
double tetrahedron can be seen in Figures 3 and 4: being one functional
and the other a 100% symmetrical geometry (with its Stella Octangula
net shown in [10]). |
| |
| The double genetic code was ‘physically’ compressed as represented
in Figures 3 and 4, indicating that through the 3-D geometry of a Stella
Octangula it’s possible to distinguish between the start amino acid
Methionine, and the non-start Methionine [10]. |
| |
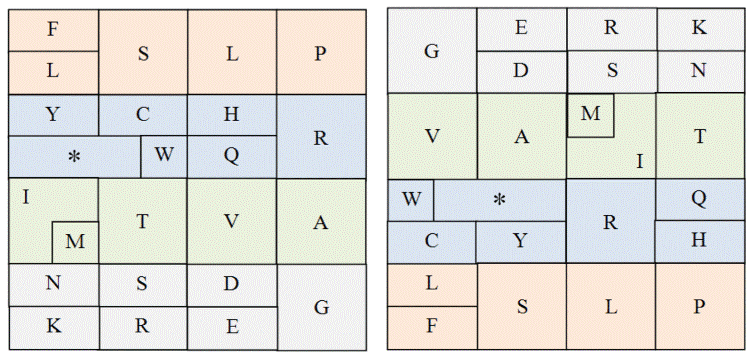
| A complex aspect of the genetic code thus far explored is the binary
correlation between the three Cartesian combinations of nucleotides (hydrogen bonds, rings, and tautomerism) by using the millenary
classical or original table of the I Ching [10], shown here compressed by
groups in Figures 5 and 6. |
| |
| Methods |
| |
| We saw in the introduction the folded Stella Octangula, as well as
its inner geometries derived from the computational analogy of the file
compression or zipping; however, we are interested here in its expansion
through the binary I Ching representation, to obtain, not only a cubical
macro representation of the genetic code, integrated by 64×6=384
codons, but also its corresponding Sphered-Cube, a resemblance of the
Cubed-Sphere widely used in Mathematics, Climatology, Geology, and
Astronomy. The practical purpose of this is two-fold: First, to provide
educational mnemonic devices; and second, to provide a useful core
for molecular modeling in bioinformatics through a, not only highly
informative, but also visually aesthetic bioinformatics tool to compare sequences of men and/or of other organisms; i.e., for the early detection
of diseases, and/or to discover genomic compatibilities between related
varieties of organisms currently misclassified as belonging to different
species, or even to a different genus, instead of what they are in reality:
varieties or sub-species, and vice versa. The geometries and binary
structures used here are: |
| |
| 1) The net or pattern to obtain a folded Stella Octangula: http://web.archive.org/web/20101124045835/ http://korthalsaltes.com/model.php?name_en=stella%20octangula, and its hidden, internal
Octahedron: http://liveweb.archive.org/ http://mathworld.wolfram.com/Octahedron.html; |
| |
| 2) The original I Ching representation attributed to Fu-Xi: http://liveweb.archive.org/ http://www.laetusinpraesens.org/musings/images/unknow_files/fuxi.jpg, |
| |
| 3) The inner cubic 3-D space for Figure 22, obtained by using
the freely available software located at: http://web.archive.org/web/20120625122500/ http://www.doka.ch/Excel3Dscatterplot.htm |
| |
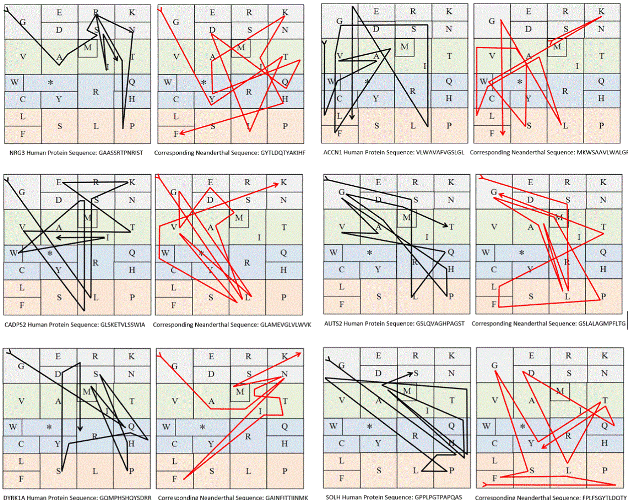
| Figures 25 and 26 show a practical application of the genetic code’s I Ching by using both the GenBank and BlastP to compare expressed
sequences and a promoter between humans and a Neanderthal, as
described there. |
| |
| Results |
| |
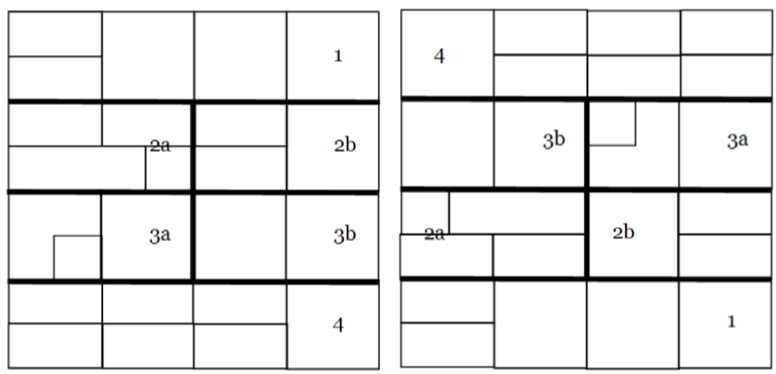
| The next figures will be devoted to show the file compression and
file expansion of the genetic code while advancing towards its macro
representation. Figures 7 and 8 show the ‘file compression’, or zipping
(zip), of codons according to their corresponding codified amino acids
or functions. |
| |
| If we now take the same groups shown in Figure 7 while removing
their colours as a previous step before exhibiting their arrows that will be
displaying the genetic code’s Yin/Yang directionality, we obtain the Figure
8, with its numbers and letters used here as a reference for the next step. |
| |
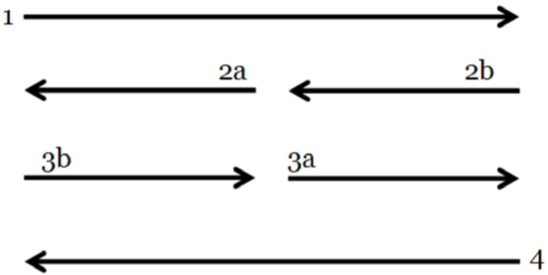
| If we now demonstrate the directionality between the groups of
codons according to their resulting amino acids or functions that are
shown in Figure 8, we obtain the Yin/Yang arrows of Figure 9. |
| |
| Figure 9 shows the directionality of the four basic components of
the Yin/Yang as derived by the ancestral I Ching: The external arrows
(flowing in opposite directions), being represented long ago by the ‘Old
Yin’ and by the ‘Old Yang’ located at its extremes, while their younger
and alternating pairing corresponds to the ‘Young Yang’ and to the
‘Young Yin’, respectively, being located at the centre while also moving
in the opposite directions. These, in general, are also representing,
both the process of DNA auto-replication (being the old strands of the
double helix of DNA in the external lines, while the new strands are the
internal), but also representing the process of DNA transcription into
smaller segments of RNA. |
| |
| If we now add the third Cartesian correlation, represented as well
by the original symbols of the I Ching, we obtain what we see in Figure
10; and, if we go one step further and we add the arrows previously seen looking at two of its contiguous walls while standing on its third side,
so we obtain Figure 11. |
| |
| The potential for the file compression within the genetic code
that we see in the preceding images can be further explored, showing
here some of its numerous possibilities: In Figure 12 we see its zipped
configuration in a circle while in Figure 13 we see such compression
both in a tetrahedron and in a cube; these images are only some
examples of the file compression applied to the genetic code analysis,
which will potentially increase the speed of the analysis of sequences of
DNA, RNA, or of proteins. If we now move one step further, we will
be able to represent every possible binary combination of the genetic
code in the classic I Ching (being only six possibilities available) within
a cube, indicating that in addition to the three Cartesian correlated
representations of the genetic code (in Appendixes. A we can see
their common points), their three reciprocals are also included (see
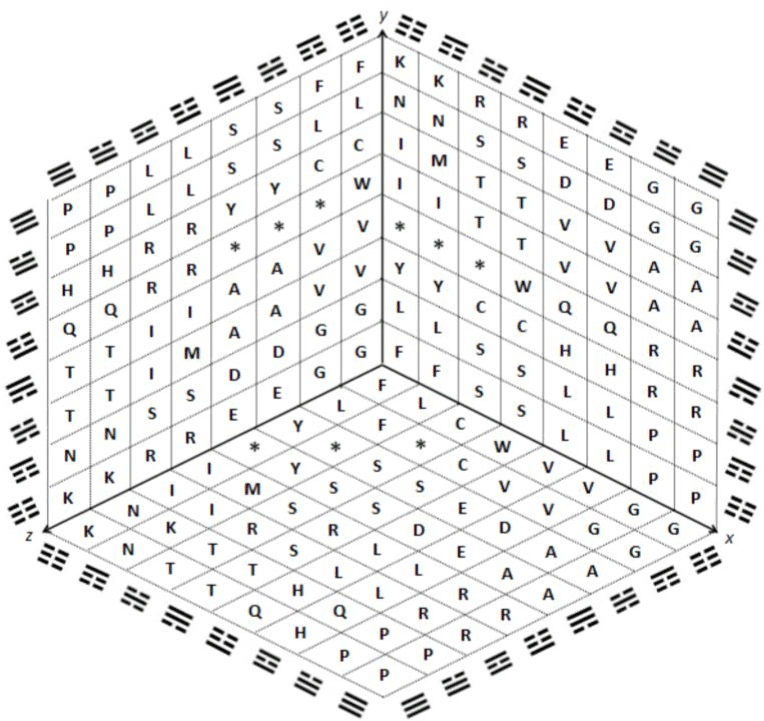
Appendixes. B); next, we will see the macro-representation of the
genetic code in 6X as derived from the I Ching, either by codons (Figure
14), by amino acids (Figure 15), or by those colored codon groups that
were shown before, but with no letters (Figure 16). |
| |
| Increasing the level of complexity for our mentioned point of
view as if standing inside the resulting cube, looking now to its two
contiguous walls, and to its floor as if these were the three standard or
normal correlations within the context shown by Figures 5, 6 and 10,
we have the resulting Figure 17 for the codons, and Figure 18 for their
resulting amino acids. |
| |
| If we now look to the reciprocals of the previously defined as standard
Cartesian I Ching genetic code correlations, we obtain Figures 19, 20, and
Appendix B. In Appendix C we see the coordinates used to obtain those
standard three correlations shown in Figures 17 and 18, as well as their
respective reciprocals shown in Figures 19 and 20, within the pair of 3-D
vector Cartesian graphics (axis x, y, z) contained in the cube. The final
product of this article will be a Sphered Cube for the genetic code (Figures
21 and 22). |
| |
| If we now correlate the relative direction of the groups of codons
according to their amino acids and/or functions, we obtain Figure 21 (for
the standard correlation), and Figure 22 (for both the standard and the
reciprocal correlations); and again, for these 3-D spatial versions were
used, both the three standard correlations (Figure 21), and the reciprocals
(Figure 22), of the Yin/Yang arrows shown in Figures 9 and 11. |
| |
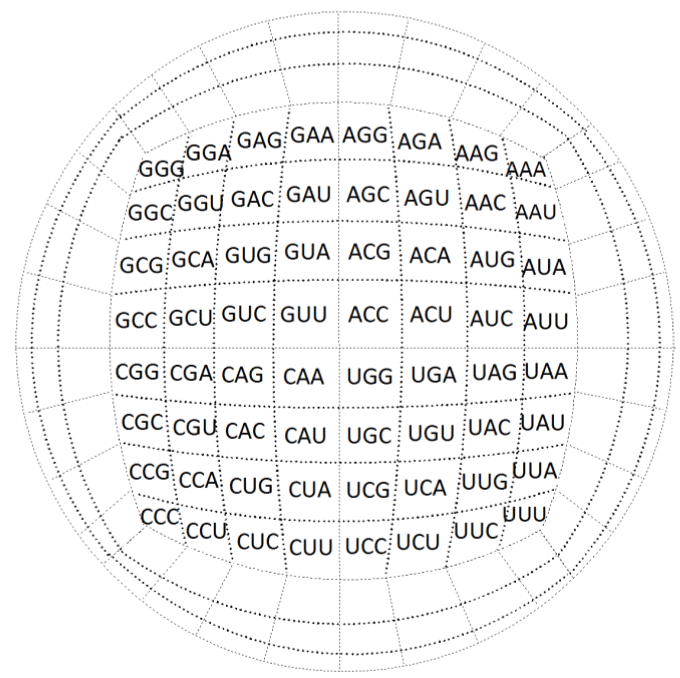
| Next, we are going to perform the ‘sphering’ of our genetic code’s
macro-cube to obtain our first meaningful spherical representation of
it, both by codons (Figure 23), and amino acids (Figure 24). |
| |
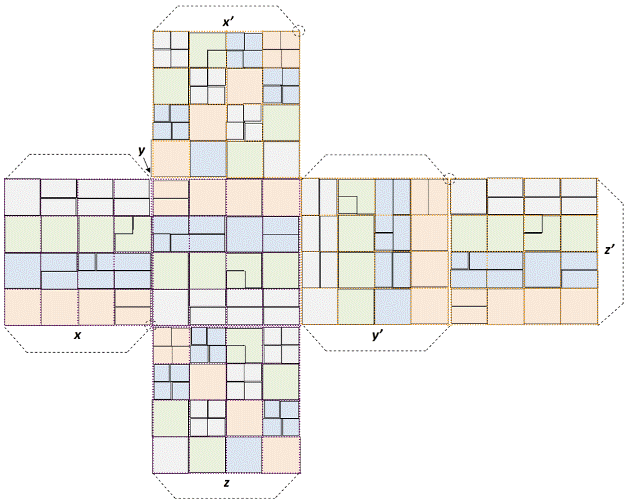
| Appendixes D show the net without words necessary to obtain the
8×8×6=384 genetic code’s cube. |
| |
| When we obtained the Sphered Cube by transforming our genetic
code’s macro-cube into a sphere, we were surprised to realize that its
resulting product was similar to the inverse reasoning of the ‘cubedsphere’,
used initially in climatology for the subdivision of the earth
[15-17], then in mechanics [18], in computational astrophysics [19,20], in mathematics [21], physics, chemistry, hydrology [22,23],
climatology, geology, meteorology [24], etc.; however, this article
seems to be the first time that such a structure has been originated
through an opposite reasoning: the ‘Sphering’ of the cube instead of
the ‘Cubing’ of a sphere, being here applied to the genetic code and to
biology in general; an early model was built to represent our planet by
using a smaller, 6×6=36 grids times six [15], and a similar image to the
one obtained here (8×8=64), but by using an inverse reasoning, was
obtained by [24, in its p. 840]. |
| |
| The keen observations done earlier by shCherbak, who has worked
several aspects of the mathematical balance of the genetic code, are
that: “…one may assume that natural computing can exist as well… such
a computing could be essence of exact gene processing and scrambling…
If that is the case, some cell organelles should work as biocomputers… the
genetic code is connected more closely to abstract notions of arithmetic
than with notions of physics or chemistry” [25]. Through the findings
presented in this article, these statements of shCherbak have been
corroborated. |
| |
| And, in a similar way to what we have seen in our introduction
when presenting the word ‘code’, shCherbak earlier equated the stop
codons with zero (Arabic, Mayan, and Computational), deeming the
zero as the supreme arithmetical abstraction and concluding that its
use by any alphabet, including the genetic code, is an indicator of what
he called ‘artificiality’ (being such artificiality in this case, the intelligent
design and purpose of the genetic code: to bring and to sustain life),
while affirming that the decimalization of the genetic code could be a
special case of the general computational power of genomics and its
molecular machinery, observing that the main reason for the origin
of a numerical system is to do mathematical calculations; shCherbak
concluded that: “there is no way to write or read any gene when no code
is available” [26].4 |
| |
| As we mentioned earlier, the Cube and its Sphered Cube presented
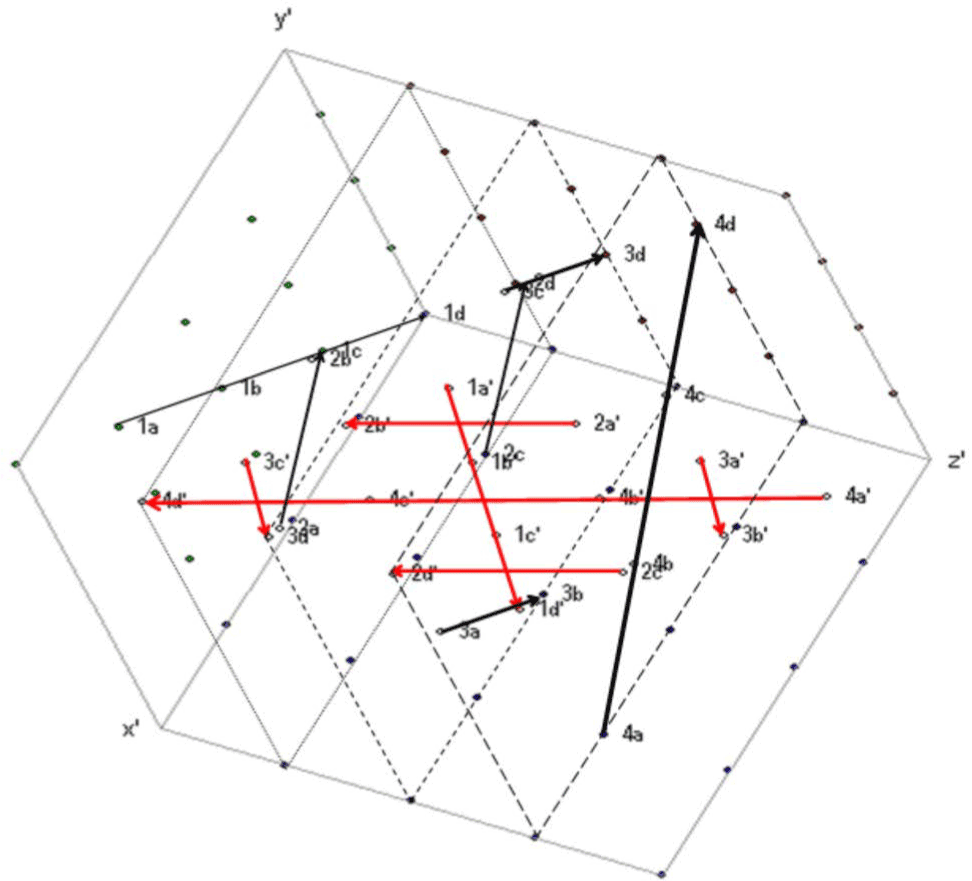
in this article can be used to compare sequences. Prompted by a reviewer and to demonstrate a practical application of our discovery, we will use
here a compressed way of it by grouping its codons according to its
resulting amino acids (Figure 25), where the matching sequenced genes
can simultaneously run gene by gene within two cubes that belong to
different organisms (in our example, Humans represented by the black
arrows and Neanderthal by the red arrows), while pointing out to their
differences by a resulting drawing. |
| |
| More recently, while I was ready to resubmit this article, it was
found that the patterns of expression are also very different between
humans and neanderthals, highly differing in their methylation and
deamination, a finding that may help explain why the expressed
sequences of humans (the human amino acids) were extremely different
(as shown in Figure 25), even corresponding, perhaps, to totally
different proteins, when compared to neanderthals (the Neanderthal’s
amino acids). As an exercise, Figure 26 shows the differences on methylation/deamination between humans and ‘a proxy’ Neanderthal
[36]. |
| |
| And, due to the fact that we are dealing with ancient sources of
information, such as the I Ching and the Neanderthal’s genome, in
Appendix E I dare to present what ancient manuscripts in Hebrew
(shown in the introduction), Aramaic and Greek, that are contained
within the Bible have to say in relation to it. Otherwise, in order to
analyze more in detail the currently full Neanderthal and Human
sequences freely available through the Max Planck institute5, I will
require of a greater computer power and memory space, as well as the
development of a computational program with the Sphered Cube as
its engine; for example, I performed earlier an unpublished analysis of
some of the differences in the mitochondrial DNA between Humans
and Neanderthal,6 however, the current study will be more graphically
appealing. |
| |
| Elsewhere, an example of a human polymorphism [9], and the
comparison of the human genome with other organisms has been presented [9,37], as well as one molecular application of the genetic
code in 3-D to differentiate the start from non-start Methionines
within proteins [10]. |
| |
| Also, and apart of exploring the 3-D properties of the genetic
code, cell cultures in 3-D have been recently used by the author of this
article in an attempt to simulate the current conditions of living tissues
through the modeling of the origin and attenuation of atherosclerosis
[38] in human osteogenic trans-differentiating vascular smooth
muscle cells (VSMCs). We saw that Lyso-phosphatidylcholine (LPC) at
physiological concentrations (10 nm), instead of the entire Low Density
Lipoprotein (LDL) or ‘bad cholesterol’, seemed to have been the main
responsible for the development of calcification in atherosclerosis [39-41], being this a defensive response of the arterial muscular cells against
the detergent-like effects of the LPC, a mechanism for those cells to
protect themselves. |
| |
| The optimization of the 3-D cell culture methodology for
atherosclerosis required more than two years; however, reference [42],
not only added the LPC that was successfully demonstrated by us [39],
while using my methodologies and materials with no credits, but also
ruined our experimental model by adding two extra, non-necessary
and non-physiological reagents: β-glycerophosphate (7.5 mM) and
ascorbic acid (50 μg/ml), being these additions reflected in their work
by the retention of hydroxyapatite within the VSMCs of their 3-D
cluster (seen at the end of their third composite illustration), instead
of being normally secreted, as it was shown in all of our Figures [38]. |
| |
| Our original work [38] demonstrates that a 3-D perspective is
highly useful in biomedical research, not only for its 3-D cell cultures,
but also for the current study of the genetic code in 3-D as shown here. |
| |
| Tidjani Negadi, a dedicated researcher of the genetic code, has been
able to find numerous instances of the number 384 within the genetic
code, a number here obtained through our I Ching cube (64×6=384);
Negadi presents at least 15 calculations to obtain the number 384
within the genetic code: |
| |
| 1) 360+tau (360)=360+24=384 [43] (number equal to the number
of atoms in the 20 amino acids, side-chain and block [44], as well as the
total number of nucleotides in the 64 RNA-codons and in the 64 DNA
codons [45]), |
| |
| 2) (73+11) +300=84+300=384 [43], |
| |
| 3) (99+74+7)+(73+41+11+20+35)+σ(360)=180+204=384 [43], |
| |
| 4) 360+(5+5+5)+9=384 [43], |
| |
| 5) 2x192=384 [43,45], |
| |
| 6) (3412/2x)-1255-67=384 (from the irregular L- and D- (integer)
tetrahedron, “a warehouse of biological information”) [43], |
| |
| 7) 192+192=384 [45] (The mean of the four pairs equal 192, the
total number of nucleobases in the 64 codons, but also the total number
of codons in DNA and RNA (2×64) [46]), |
| |
| 8) 768/2=384 [46], |
| |
| 9) phi(2)(psi)=384 [46], |
| |
| 10) psi upper bar(0,1,0,0)(psi)=psi upper bar(0,0,3,0)(psi)=384 [46], |
| |
| 11) Writing 3276+676 and introducing the identity “-384+384”
(384=384 or “384-384=0”), we get 2892+1060 which is the “partition
of nucleon-numbers between the four-codon set and the non-four codon
set” [46], |
| |
| 12) phi(1152)=384 [46], |
| |
| 13) 313+71=384=204+(9x20), |
| |
| 14) Filatov [47] also “established a nucleon number balance in
two pairs of faces 628+627=626+629=1255. Now computing the atom
number in the amino acids in the two pairs of faces, we find 108+97=205
and 104+99=203 (without the side-chains) or 198+187=385 and
194+189=383 (including the side-chains). These are near balances and
the average of the two pairs gives 204 and 384 atoms, respectively, for
the correct atom number. Now from our numbers 139 (34th prime) and
1304, we could take the sum of the prime factors and the prime indices to get 173 and 210 with sum 383. This is the nucleon number in one
of the two pairs of faces mentioned above …and we could reach 384
by just adding the Omega-function of the prime number 139 which is
simply 1. Also, by re-arranging the terms we have (139+34+1+6)=180
and (163+38+3)=204. The first number is the total atom number in
the 20 blocks and second is the atom number in the 20 side-chains”. So,
concluding the last computation, we have (139+34+1+6)+(163+38+3
)=384=180+204 [The points 13 and 14 were found in Negadi’s blog]. |
| |
| 15) Plus, a music related one: “The numbers 192 and 384 seem very
important numbers” [Plato did choose 192 as a starting point while
Timaeus the Locrian did choose 384] “…they are basic numbers in the
Pythagorean music tuning system” [45] (also see [44]). |
| |
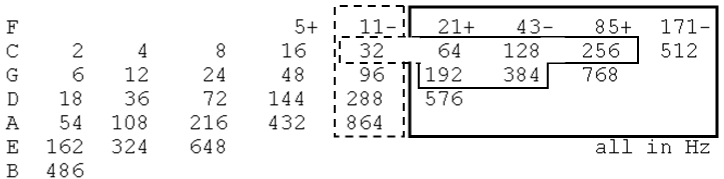
| Figure 27 shows how both our 384-grid Cube and Sphered Cube
representation of the genetic code can be compared to the Pythagorean
music series. |
| |
| Using only proportions of 2 and 3 (± mean by 1/3, in row F),
the frequency relationships seen in Figure 27, when applied to the
building-up of the I Ching genetic code are: 64, 128, 192, 256, and 384;
however, the I Ching’s successive number 320 that appears between 256
and 384 is not present; the numbers that are actually present, in the
current context are: 64×1=64, 64×2=128, 64×3=192, 64×4=256, [the
next consecutive one, 64×5=320, is absent in the Pythagorean music
system], and 64×6=384. Additional multiples of 64 that are seen in the
Pythagorean musical scale are: 64×8=512, 64×9=576 and 64×12=768. |
| |
| Notably, if the numbers within the left side column (headed by 11-
in the row F) are multiplied by 10, all of them became also multiples
of 64, including the previously missing 64×5=320, plus 64×15=960,
64×45=2880, and 64×135=8640. |
| |
| Regarding such discrepancy seen in lines C and E (and even B) of the
Pythagorean music series, Ray Tomes observes: “C is incorrectly related
to E and B… as Galilei observed, C and E want to be in the proportion 4:5
(which is 64:80) and we have 64:81 so it just misses… The equitempered
scale is designed around the ratio 2 because each semitone is a ratio of
1 to the twelfth root of 2. As it turns out this accommodates ratios of 3
almost perfectly, which is of course why it was chosen… I invented a
system which I call AJI for Automatic Just Intonation. I realised [sic] that
an electronic keyboard doesn’t have to make compromises when selecting
frequencies as there is no reason that a single key cannot produce
different frequencies depending on the circumstances…” In our case, as
mentioned earlier, we multiplied by ten the whole column 11- to make
it ‘fit’ the pattern of our design, and it worked, providing us with an
artificially ‘perfect’ symmetry, as seen in Figure 27. |
| |
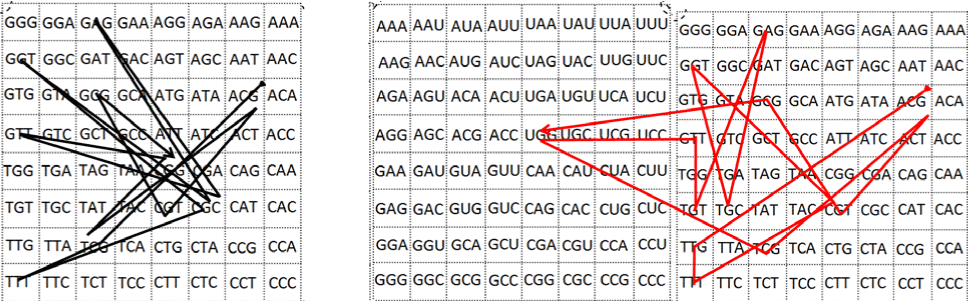
| 16) In our case, the number 384 was also obtained by counting the
individual binary components within any of the I Ching’s genetic code
tables (Figures 5, 6, 10), where we see that each cell is composed by a
trigram from the y axis, over which a trigram from the x axis is added,
integrating an hexagram, so when multiplying one hexagram by the
totality of the cells, we also obtain the number 384: 6×64=384. |
| |
| Finally, and as the last project of my free programming edX Harvard
University class (cs50x), I decided to put motion to my earlier work to
illustrate the complementary pairing by rows of the basic structure of
the Figure 5 of this article, which is similar, but not identical, to the
first Table (T.1) of Figure 1 of my previously related publication [10].
The project in its final form can be seen at: http://www.youtube.com/watch?v=0aacCShsmWo, coupled to an explanation of it. The basic 8 sprites and 9 scripts used to develop the animation for this video can be
downloaded at its M.I.T. ‘source code’: |
| |
| http://scratch.mit.edu/projects/fdocc/3226378 Our current
experimental lab work dealing with 3-D cubic cylinders and crystallized
geometries can also be seen at: http://youtu.be/oYd96rj_51Q |
| |
| Conclusion |
| |
| Biodiversity on earth is based on the flexibility and plasticity of the
genetic code. This is just the start of studies demonstrating the enormous
plasticity of the genetic code, being its current representations and its
future possible ones like a metaphorical reflection of the real plasticity
contained within living organisms [10], being this the reason for their
great variability and adaptation as seen in the exuberant biodiversity
that surrounds us. |
| |
| We also showed here the reverse engineering in 2-D of a previously
obtained functional tetrahedron in 3-D [10]. The resulting products
were a square (Figure 1) and a circle (Figure 2). We also folded a 3-D
double genetic code representation composed by a Stella Octangula
(Figsures 3 and 4) found earlier [10]. |
| |
| The rotation seen in Figure 9 from row 1 to row 4 can also be
perceived as evocative of the actual helical rotation that is seen in the
double helix [14]. The two circular and compressed representations
of the genetic code with the stop codon at the centre that are shown
in Figure 12, if they were rotating through a pin at their centre, while
an external arrow were pointing at their respective codons, if it were
stopped at a line between two of these portions, it will go then to the
stop codons located at its centre. |
| |
| The inclined rotational axis of 45 degrees shown in Figures 14-16
(represented there by the intermittent circles, their net can be seen in
Appendix D), remind us of the inclined orbital axis of the earth; these
diagonals of computational geometry within a cube are deemed to be
located either as a π/4 or a 3π/4, depending if the relative view is from
left to right or vice versa [48-50] shows the software used to develop
(Figure 22). |
| |
| The zipping or compression of the codons by groups, both in the
3-D and in 2-D representations of the genetic code discussed here
may increase the speed of our comparison of sequences, as well as our
explorations of the genetic code within curved spaces, spaces studied
in my classes related to the foundations of computer graphics;7 i.e., the
resulting cubed-sphere may be useful to compare sequences, either from
wildlife or from humans in an appealing visually graphic way, noticing
also that the file expansion shown here is not the same as the unzipping
of the computer files, being this the product of the restoration to the
same size of the original file as it was before, while the file expansion
shown here results in the increase of informational content. |
| |
| Here it was also shown that human genes translated from the
brain and from a promoter in osteoblasts, are compared to their
corresponding Neanderthal sequences to show their differences and to
demonstrate the use of the I Ching’s genetic code in 3-D (Figures 25 and
26). After these findings, we can declare with confidence that the oldest
known representation of the Genetic Code is the binary I Ching, also
known as the Book of Changes or Book of Mutations, corresponding
these changes to biological changes or mutations, a book preserved
by the Chinese, but evidently being older than them, a survivor of the
burning of ancient documents by the Emperor Chi (also known as Qi). |
| |
| In relation to this and to my previous works, a recent article
declared: “Those two properties (ie, symmetry and periodicity) act as the harmony between the chosen geometry and the biological reality.
Graphical representation of DNA sequences based on mono, di,
trinucleotides, etc. need to consider this harmony. Otherwise, it would
merely be an instance of displaying the nucleotides (eg, mononucleotide,
dinucleotide, codon) which have little biological sense” [51]; with the
current work, the corroboration of such observations is advanced. |
| |
| A recent article [52], making also reference to one of my earliest
works related on the 2-D circular genetic code, when alluding to “the
palindromes of the genetic code”, declared: “that some researchers have
tried to restrict at peculiar symmetries”; my answer is that in order to
learn from the continuous wisdom manifested within the genetic code,
certainly the less limited our analyses are, the best their output will be; I
strive to encourage every student and researcher of the genetic code to
explore the fullness of its possibilities without dismay. |
| |
| Once more, the enormous plasticity that we see in the study of the
genetic code is a reflection of the real plasticity contained within living
organisms, being this the ultimate reason for their great variability and
adaptation. The great variability seen in nature can better be understood
by studying its source, the genetic code; for example, in my earlier studies
[8,9] of the 2-D classic circular genetic code [5], I recently realized that
the Hydroxyl amino acids, those ones that are phosphorilatable in order
to activate the dormant enzymes, are also located in a balancing way
within quadrant 1 (4 Ser+2 Tyr), and quadrant 3 (2 Ser+4 Thr). |
| |
| The use of the Yin/Yang (Yin and Yang) in this article is within
the context of the Genetic Code in 3-D, something apparently known
and fully represented since a long time; i.e., in the I Ching or Book of
Changes; however, the use of this concept is not new within medicine
and molecular biology; a few set of examples from the numerous found
within PubMed (currently ~500 references), are given at [53-85]. |
| |
| Here, by grouping the reiterations or “redundancies” of the genetic
code, reiterations apparently designed to preserve its functionality
while allowing the adaptability of living organisms, a more detailed
picture appeared, both in 2-D and in 3-D, when comparing the
Cartesian binary combinations of the properties of nucleotides, then
the directions of the Yin and of the Yang arrows suddenly appeared. It is
precisely this reiterated redundancy of the genetic code, the basis for its
‘file compression’ in bioinformatics and biodiversity; additionally, it was
shown that the number 384, obtained in this article by multiplying 64x6
within all the possible binary comparisons of the genetic code derived
from the I Ching, has also been found within the mathematics derived
from the molecules that integrate the genetic code, as seen above when
our current work is compared with that of Negadi [43,45-46], and
others [44,47]. |
| |
| On concluding, I may say that the main findings of this work are:
That the Yin/Yang’s genetic code directions are similar to those that are
presented by the actual separate DNA strands in relation to their autoreplicating
new strands, and/or in relation to their new RNA products,
specially by the plus strand at the moment of transcription for the last
example. Also including the minus strand, and its smaller replicating
segments known as the Okazaki fragments for the DNA autoreplication.
The findings presented here can help in bioinformatics;
i.e., by locating at the center of a computer program the genetic code
in its cubic or spherical form, not only to compare sequences in an
attractive or aesthetic way, but also to provide didactic illustrations of
these proceedings. |
| |
| Acknowledgements |
| |
| I thank God for giving me the time and strength to do this work, and to my wife Tracy L. Duncan for her unconditional support; to Reid and Sheryl McNally,
Stephen and Brenda Sparks, Cathy Keener, Eliezer Mendelssohn, Fernando
Paularena, Glenn Lewis, Lynn D. Wills, David J. Lewis, Margina Huq, Sheldon
Yoder, Bruno Bazan, Zachary Greenbauer, Chris Caldwell, Daniel Garcia, Mark
Skipski, Maria Lantsman, Olivia Nguyen, and to the rest of my coworkers that make
all the daily work enjoyable; and also to my parents, sisters, nephews, to the inlaws,
and to my students Ana Baleva and Karen Scarbrough, as well as to my
spiritual and physical families and friends. This manuscript was supported in part
by the NIH grant T32 HL-07812. |
| |
| 1 http://web.archive.org/web/20101129051637/ http://rumkin.com/tools/cipher/
numbers.php |
| |
| 2 Abbr.: Amino acids: A: alanine (Ala), V: valine (Val), I: isoleucine (Ile), L: leucine
(Leu), M: methionine (Met), F: phenylalanine (Phe), W: tryptophan (Trp), D: aspartic
acid (Asp), N: asparagine (Asn), E: glutamic acid (Glu), Q: glutamine (Gln), R:
arginine (Arg), K: lysine (Lys), S: serine (Ser), T: threonine (Thr), G: glycine (Gly),
P: proline (Pro), H: histidine (His), C: cysteine (Cys), Y: tyrosine (Tyr). Nucleotides
(nt): U: uracil, C: cytosine, A: adenine, G: guanine, T: thymine. n: any nt, r: purines,
y: pyrimidines. |
| |
| 3 Psalm 139:16, http://biblehub.com/interlinear/psalms/139-16.htm. |
| |
| 4 Its mdi file is available at: http://liveweb.archive.org/ http://tinyurl.com/shcherbak |
| |
| 5 http://cdna.eva.mpg.de/neandertal |
| |
| 6 http://fdocc.ucoz.com/index/0-121 |
| |
| 7 Class CS184.1x Foundations of Computer Graphics, imparted by Ramamoorthi,
from U.C. Berkeley (free at edX). |
| |
|
| References |
| |
- Bishop D (2003) Introduction to cryptography with Java applets. Jones & Bartlett Learning 16-17.
- Leonhard W (2013) Windows 8.1 All-in-One for Dummies.Wiley, 1056.
- Nirenberg MW (1965) 64 triplets and complementary triplets. Unpublished laboratory note.
- Crick FH (1967) The Croonian lecture, 1966. The genetic code. Proc R Soc Lond B Biol Sci 167: 331-347.
- Bresch C, Hausmann R (1972) Der genetische code. In: Bresch C, Hausmann R, eds. Klassische und Molekulare Genetik. Springer (Berlin) 243–278.
- Fujiomoto M (1987) Tetrahedral codon stereo-table. U.S. Patent 4,702,704, 9 p.
- Keppleri I (1619) Harmonices mundi. Johannes Plancus (Austria).
- Castro-Chavez F (2010) The rules of variation: amino acid exchange according to the rotating circular genetic code. J Theor Biol 264: 711-721.
- Castro-Chavez F (2012) The rules of variation expanded, implications for the research on compatible genomics. Biosemiotics 5:121-145.
- Castro-Chavez F (2012) Defragged binary I Ching genetic code chromosomes compared to Nirenberg’s and transformed into rotating 2D circles and squares and into a 3D 100% symmetrical tetrahedron coupled to a functional one to discern start from non-start methionines through a stella octangula. JPSCB 1 : 1-24.
- Castro-Chavez F (2012) A Tetrahedral Representation of the Genetic Code Emphasizing Aspects of Symmetry. BIOcomplexity 2012: 1-6.
- Castro-Chavez F (2011) The Quantum Workings of the Rotating 64-Grid Genetic Code. Neuroquantology 9: 728-746.
- Swanson R (1984) A unifying concept for the amino acid code. Bull Math Biol 46: 187-203.
- WATSON JD, CRICK FH (1953) Molecular structure of nucleic acids; a structure for deoxyribose nucleic acid. Nature 171: 737-738.
- Sadourny R (1972) Conservative finite-difference approximations of the primitive equations on quasi-uniform spherical grids. Mon Weather Rev 100:136–144.
- Rancic M, Purser J, Messinger F (1996) A global shallow water model using an expanded spherical cube: Gnomonic versus conformal coordinates. Q J Roy Meteor Soc 122:959–982.
- Purser J, Rancic M (1998) Smooth quasi-homogeneous gridding of the sphere. Q J Roy Meteor Soc 124:637–647.
- Koldoba AV, Romanova MM, Ustyugova GV, Lovelace RVE (2002) Three-dimensional magnetohydrodynamic simulations of accretion to an inclined rotator: The "cubed sphere" method. ApJ 576 : L53-L56.
- Romanova MM, Ustyugova GV, Koldoba AV, Wick JV, Lovelace RVE (2003) Three-dimensional simulations of disk accretion to an inclined dipole. I. Magnetospheric flows at different T. ApJ 595 : 1009–1031.
- Fragile PC, Lindner CC, Anninos P, Salmonson JD (2009) Application of the cubed-sphere grid to tilted black hole accretion disks. ApJ 691:482–494.
- Ronchi C, Iacono R, Paolucci P (1996) The ‘‘Cubed-Sphere:’’ a new method for the solution of partial differential equations in spherical geometry. J Computat Phys 124 : 93-114.
- Cheruvua V, Nairb RD, Tufoa HM (2007) A spectral finite volume transport scheme on the cubed-sphere. Appl Numer Math 57 :1021–1032.
- Chen CG, Xiao F, Li XL, Yang Y (2011) A multi-moment transport model on cubed-sphere grid. Int J Numer Meth Fl 67 : 1993–2014.
- Washington WM1, Buja L, Craig A (2009) The computational future for climate and Earth system models: on the path to petaflop and beyond. Philos Trans A Math Phys Eng Sci 367: 833-846.
- shCherbak VI (2003) Arithmetic inside the universal genetic code. Biosystems 70: 187-209.
- shCherbak V (2008) The arithmetical origin of the genetic code. In: Barbieri, Marcello (Ed.) The Codes of Life. Springer (Berlin). 153–188.
- Strausberg RL1, Feingold EA, Grouse LH, Derge JG, Klausner RD, et al. (2002) Generation and initial analysis of more than 15,000 full-length human and mouse cDNA sequences. Proc Natl Acad Sci U S A 99: 16899-16903.
- Gregersen N1, Dahl HA, Buttenschřn HN, Nyegaard M, Hedemand A, et al. (2012) A genome-wide study of panic disorder suggests the amiloride-sensitive cation channel 1 as a candidate gene. Eur J Hum Genet 20: 84-90.
- Beunders G1, Voorhoeve E, Golzio C, Pardo LM, Rosenfeld JA, et al. (2013) Exonic deletions in AUTS2 cause a syndromic form of intellectual disability and suggest a critical role for the C terminus. Am J Hum Genet 92: 210-220.
- Guimerá J1, Casas C, Pucharcňs C, Solans A, Domčnech A, et al. (1996) A human homologue of Drosophila minibrain (MNB) is expressed in the neuronal regions affected in Down syndrome and maps to the critical region. Hum Mol Genet 5: 1305-1310.
- Kamei M, Webb GC, Young IG, Campbell HD (1998) SOLH, a human homologue of the Drosophila melanogaster small optic lobes gene is a member of the calpain and zinc-finger gene families and maps to human chromosome 16p13.3 near CATM (cataract with microphthalmia). Genomics 51 : 197-206.
- Briggs AW, Good JM, Green RE, Krause J, Maricic T, et al. (2009) Targeted retrieval and analysis of five Neandertal mtDNA genomes. Science 325: 318-321.
- Green RE, Malaspinas AS, Krause J, Briggs AW, Johnson PL, et al. (2008) A complete Neandertal mitochondrial genome sequence determined by high-throughput sequencing. Cell 134: 416-426.
- Green RE, Krause J, Briggs AW, Maricic T, Stenzel U, et al. (2010) A draft sequence of the Neandertal genome. Science 328: 710-722.
- Gibbons A (2010) Paleogenetics. Close encounters of the prehistoric kind. Science 328: 680-684.
- Gokhman D, Lavi E, Prüfer K, Fraga MF, Riancho JA, et al. (2014) Reconstructing the DNA methylation maps of the Neandertal and the Denisovan. Science 344: 523-527.
- Castro-Chavez F (2011) Most Used Codons per Amino Acid and per Genome in the Code of Man Compared to Other Organisms According to the Rotating Circular Genetic Code. Neuroquantology 9.
- Castro-Chavez F, Vickers KC, Lee JS, Tung CH, Morrisett JD (2013) Effect of Lyso-phosphatidylcholine and schnurri-3 on osteogenic transdifferentiation of vascular smooth muscle cells to calcifying vascular cells in 3D culture. Biochim Biophys Acta 1830 : 3828–3834.
- Vickers KC1, Castro-Chavez F, Morrisett JD (2010) Lyso-phosphatidylcholine induces osteogenic gene expression and phenotype in vascular smooth muscle cells. Atherosclerosis 211: 122-129.
- Castro-Chavez F, Morrisett JD (2011)Osteogenic transdifferentiation of vascular smooth muscle cells to calcifying vascular cells in 3D culture: enhancement by lyso-phosphatidylcholine and attenuation by Schnurri-3. Poster 1108-116, ACC 11, J Am Coll Cardiol 57: E1548.
- Castro-Chavez F, Lee JS, Tung CH, Morrisett JD (2011) Osteogenic transdifferentiation of vascular smooth muscle cells in 3D, detecting hydroxyapatite with a specific probe derived from the alpha region of osteocalcin, Rolanette and Berdon Lawrence Bone Disease Program of Texas Scientific Retreat, Marriott Medical Center Hotel. Baylor College of Medicine and MD Anderson Cancer Center, p3.
- Lee JS1, Morrisett JD, Tung CH (2012) Detection of hydroxyapatite in calcified cardiovascular tissues. Atherosclerosis 224: 340-347.
- Negadi T (2012) the irregular (integer) tetrahedron as a warehouse of biological information. Symm Cult Sci 23(3-4):403-426.
- Rakocevic MM (2013) Harmonic mean as a determinant of the genetic code. arXiv:1305.5103v4 [q-bio.OT], 20 p.
- Negadi T (2011) The multiplet structure of the genetic code, from one and small number. Neuroquantology 9 : 379-384.
- Negadi T (2011) A “quantum-like” approach to the genetic code. Neuroquantology 9 : 785-798.
- Filatov F (2009) A molecular mass gradient is the key parameter of the genetic code organization. arXiv:0907.3537v1 [q-bio.OT], 9 p.
- Mukkamala P, Palvolgyi D (2012) Drawing cubic graphs with the four basic slopes. Lect Notes Comput Sc (7034) 254-265.
- Jeon G, Fang Y, Lee K, Kang K, Jeong J (2012) Improved low-complexity interpolation algorithm for deinterlacing. OJEEE 2 : 259-263.
- Doka G (2002) Life Cycle Assessment of municipal solid waste incineration with the PECK technology ARGE PECK (Zurich) 80 p.
- Bari AT, Reaz MR, Islam AK, Choi HJ, Jeong BS (2013) Effective Encoding for DNA Sequence Visualization Based on Nucleotide's Ring Structure. Evol Bioinform Online 9: 251-261.
- Godina-Nava JJ (2013) Horizontal symmetry in the algebraic approach of genetic code. arXiv:1302.3289v1 [physics.gen-ph], 20 p.
- Mango R, Predazzi IM, Romeo F, Novelli G (2011) LOX-1/LOXIN: the yin/yang of atheroscleorosis. Cardiovasc Drugs Ther 25: 489-494.
- Rissman EF (2008) Roles of oestrogen receptors alpha and beta in behavioural neuroendocrinology: beyond Yin/Yang. J Neuroendocrinol 20: 873-879.
- Weber DS, Griendling KK (2003) The yin/yang of superoxide dismutase mimetics: potential cardiovascular therapies? Br J Pharmacol 139: 1059-1060.
- Mann DL (2002) The yin/yang of innate stress responses in the heart. Cold Spring Harb Symp Quant Biol 67: 363-370.
- Weissmann G, Goldstein I, Hoffstein S, Chauvet G, Robineaux R (1975) Yin/Yang modulation of lysosomal enzyme release from polymorphonuclear leukocytes by cyclic nucleotides. Ann N Y Acad Sci 256: 222-232.
- Clark CE, Liu Y, Cooper HM (2014) The Yin and Yang of Wnt/Ryk axon guidance in development and regeneration. Sci China Life Sci 57: 366-371.
- Wu XS, Wu LG (2014) The yin and yang of calcium effects on synaptic vesicle endocytosis. J Neurosci 34: 2652-2659.
- Martin DS, Grocott MP (2013) III. Oxygen therapy in anaesthesia: the yin and yang of O2. Br J Anaesth 111: 867-871.
- Olsen LF, Issinger OG, Guerra B (2013) The Yin and Yang of redox regulation. Redox Rep 18: 245-252.
- Nie J1, Han X, Shi Y (2013) SAD-A and AMPK kinases: the "yin and yang" regulators of mTORC1 signaling in pancreatic β cells. Cell Cycle 12: 3366-3369.
- Martins-Green M (2013) The Yin and Yang of Integrin Function in Re-Epithelialization During Wound Healing. Adv Wound Care (New Rochelle) 2: 75-80.
- Shiekhattar R (2013) The Yin and Yang of enhancer-like RNAs. EMBO J 32: 2533-2534.
- Han HW, Ohn JH, Moon J, Kim JH (2013) Yin and Yang of disease genes and death genes between reciprocally scale-free biological networks. Nucleic Acids Res 41: 9209-9217.
- Pradere JP, Dapito DH, Schwabe RF (2013) The Yin and Yang of Toll-like receptors in cancer. Oncogene .
- Tobin DM, Ramakrishnan L (2013) TB: the Yin and Yang of lipid mediators. Curr Opin Pharmacol 13: 641-645.
- Stange DE, Clevers H (2013) Concise review: the yin and yang of intestinal (cancer) stem cells and their progenitors. Stem Cells 31: 2287-2295.
- Edgerton DS, Cherrington AD (2013) Glucagon's yin and yang effects on hepatic glucose production. Nat Med 19: 674-675.
- Axelrad MD, Budagov T, Atzmon G (2013) Telomere length and telomerase activity; a Yin and Yang of cell senescence. J Vis Exp : e50246.
- Munro CA (2013) Chitin and glucan, the yin and yang of the fungal cell wall, implications for antifungal drug discovery and therapy. Adv Appl Microbiol 83: 145-172.
- Alvarez AF, Rodriguez C, Georgellis D (2013) Ubiquinone and menaquinone electron carriers represent the yin and yang in the redox regulation of the ArcB sensor kinase. J Bacteriol 195: 3054-3061.
- Tan M, Wang M, Zhou XL, Yan W, Eriani G, et al. (2013) The Yin and Yang of tRNA: proper binding of acceptor end determines the catalytic balance of editing and aminoacylation. Nucleic Acids Res 41: 5513-5523.
- So AY, Zhao JL, Baltimore D (2013) The Yin and Yang of microRNAs: leukemia and immunity. Immunol Rev 253: 129-145.
- Jiménez J, Truman AW, Menoyo S, Kron SJ, Clotet J (2013) The yin and yang of cyclin control by nutrients. Cell Cycle 12: 865-866.
- Zhang EY, Kong KF, Altman A (2013) The yin and yang of protein kinase C-theta (PKCθ): a novel drug target for selective immunosuppression. Adv Pharmacol 66: 267-312.
- Burke AJ, Sullivan FJ, Giles FJ, Glynn SA (2013) The yin and yang of nitric oxide in cancer progression. Carcinogenesis 34: 503-512.
- Vij N, Downey GP (2013) The yin and yang of cystic fibrosis transmembrane conductance regulator function: implications for chronic lung disease. Am J Respir Crit Care Med 187: 120-122.
- Vasquez KM, Wang G (2013) The yin and yang of repair mechanisms in DNA structure-induced genetic instability. Mutat Res 743-744: 118-31.
- Kubli DA, Gustafsson ĹB (2012) Mitochondria and mitophagy: the yin and yang of cell death control. Circ Res 111: 1208-1221.
- Weinberg A1, Jin G, Sieg S, McCormick TS (2012) The yin and yang of human Beta-defensins in health and disease. Front Immunol 3: 294.
- Hein GJ, Baker C, Hsieh J, Farr S, Adeli K (2013) GLP-1 and GLP-2 as yin and yang of intestinal lipoprotein production: evidence for predominance of GLP-2-stimulated postprandial lipemia in normal and insulin-resistant states. Diabetes 62 : 373-381.
- Adams EJ, Luoma AM (2012) The yin and yang of CD1d recognition. Nat Immunol 13: 814-815.
- Sladek FM (2012) The yin and yang of proliferation and differentiation: cyclin D1 inhibits differentiation factors ChREBP and HNF4α. Cell Cycle 11: 3156-3157.
- Hou J (2012) The yin and yang of claudin-14 function in human diseases. Ann N Y Acad Sci 1258: 185-190.
|
| |
References
|
- Bishop D (2003) Introduction to cryptography with Java applets. Jones & Bartlett Learning 16-17.
- Leonhard W (2013) Windows 8.1 All-in-One for Dummies.Wiley, 1056.
- Nirenberg MW (1965) 64 triplets and complementary triplets. Unpublished laboratory note.
- Crick FH (1967) The Croonian lecture, 1966. The genetic code. Proc R Soc Lond B Biol Sci 167: 331-347.
- Bresch C, Hausmann R (1972) Der genetische code. In: Bresch C, Hausmann R, eds. Klassische und Molekulare Genetik. Springer (Berlin) 243–278.
- Fujiomoto M (1987) Tetrahedral codon stereo-table. U.S. Patent 4,702,704, 9 p.
- Keppleri I (1619) Harmonices mundi. Johannes Plancus (Austria).
- Castro-Chavez F (2010) The rules of variation: amino acid exchange according to the rotating circular genetic code. J Theor Biol 264: 711-721.
- Castro-Chavez F (2012) The rules of variation expanded, implications for the research on compatible genomics. Biosemiotics 5:121-145.
- Castro-Chavez F (2012) Defragged binary I Ching genetic code chromosomes compared to Nirenberg’s and transformed into rotating 2D circles and squares and into a 3D 100% symmetrical tetrahedron coupled to a functional one to discern start from non-start methionines through a stella octangula. JPSCB 1 : 1-24.
- Castro-Chavez F (2012) A Tetrahedral Representation of the Genetic Code Emphasizing Aspects of Symmetry. BIOcomplexity 2012: 1-6.
- Castro-Chavez F (2011) The Quantum Workings of the Rotating 64-Grid Genetic Code. Neuroquantology 9: 728-746.
- Swanson R (1984) A unifying concept for the amino acid code. Bull Math Biol 46: 187-203.
- WATSON JD, CRICK FH (1953) Molecular structure of nucleic acids; a structure for deoxyribose nucleic acid. Nature 171: 737-738.
- Sadourny R (1972) Conservative finite-difference approximations of the primitive equations on quasi-uniform spherical grids. Mon Weather Rev 100:136–144.
- Rancic M, Purser J, Messinger F (1996) A global shallow water model using an expanded spherical cube: Gnomonic versus conformal coordinates. Q J Roy Meteor Soc 122:959–982.
- Purser J, Rancic M (1998) Smooth quasi-homogeneous gridding of the sphere. Q J Roy Meteor Soc 124:637–647.
- Koldoba AV, Romanova MM, Ustyugova GV, Lovelace RVE (2002) Three-dimensional magnetohydrodynamic simulations of accretion to an inclined rotator: The "cubed sphere" method. ApJ 576 : L53-L56.
- Romanova MM, Ustyugova GV, Koldoba AV, Wick JV, Lovelace RVE (2003) Three-dimensional simulations of disk accretion to an inclined dipole. I. Magnetospheric flows at different T. ApJ 595 : 1009–1031.
- Fragile PC, Lindner CC, Anninos P, Salmonson JD (2009) Application of the cubed-sphere grid to tilted black hole accretion disks. ApJ 691:482–494.
- Ronchi C, Iacono R, Paolucci P (1996) The ‘‘Cubed-Sphere:’’ a new method for the solution of partial differential equations in spherical geometry. J Computat Phys 124 : 93-114.
- Cheruvua V, Nairb RD, Tufoa HM (2007) A spectral finite volume transport scheme on the cubed-sphere. Appl Numer Math 57 :1021–1032.
- Chen CG, Xiao F, Li XL, Yang Y (2011) A multi-moment transport model on cubed-sphere grid. Int J Numer Meth Fl 67 : 1993–2014.
- Washington WM1, Buja L, Craig A (2009) The computational future for climate and Earth system models: on the path to petaflop and beyond. Philos Trans A Math Phys Eng Sci 367: 833-846.
- shCherbak VI (2003) Arithmetic inside the universal genetic code. Biosystems 70: 187-209.
- shCherbak V (2008) The arithmetical origin of the genetic code. In: Barbieri, Marcello (Ed.) The Codes of Life. Springer (Berlin). 153–188.
- Strausberg RL1, Feingold EA, Grouse LH, Derge JG, Klausner RD, et al. (2002) Generation and initial analysis of more than 15,000 full-length human and mouse cDNA sequences. Proc Natl Acad Sci U S A 99: 16899-16903.
- Gregersen N1, Dahl HA, Buttenschřn HN, Nyegaard M, Hedemand A, et al. (2012) A genome-wide study of panic disorder suggests the amiloride-sensitive cation channel 1 as a candidate gene. Eur J Hum Genet 20: 84-90.
- Beunders G1, Voorhoeve E, Golzio C, Pardo LM, Rosenfeld JA, et al. (2013) Exonic deletions in AUTS2 cause a syndromic form of intellectual disability and suggest a critical role for the C terminus. Am J Hum Genet 92: 210-220.
- Guimerá J1, Casas C, Pucharcňs C, Solans A, Domčnech A, et al. (1996) A human homologue of Drosophila minibrain (MNB) is expressed in the neuronal regions affected in Down syndrome and maps to the critical region. Hum Mol Genet 5: 1305-1310.
- Kamei M, Webb GC, Young IG, Campbell HD (1998) SOLH, a human homologue of the Drosophila melanogaster small optic lobes gene is a member of the calpain and zinc-finger gene families and maps to human chromosome 16p13.3 near CATM (cataract with microphthalmia). Genomics 51 : 197-206.
- Briggs AW, Good JM, Green RE, Krause J, Maricic T, et al. (2009) Targeted retrieval and analysis of five Neandertal mtDNA genomes. Science 325: 318-321.
- Green RE, Malaspinas AS, Krause J, Briggs AW, Johnson PL, et al. (2008) A complete Neandertal mitochondrial genome sequence determined by high-throughput sequencing. Cell 134: 416-426.
- Green RE, Krause J, Briggs AW, Maricic T, Stenzel U, et al. (2010) A draft sequence of the Neandertal genome. Science 328: 710-722.
- Gibbons A (2010) Paleogenetics. Close encounters of the prehistoric kind. Science 328: 680-684.
- Gokhman D, Lavi E, Prüfer K, Fraga MF, Riancho JA, et al. (2014) Reconstructing the DNA methylation maps of the Neandertal and the Denisovan. Science 344: 523-527.
- Castro-Chavez F (2011) Most Used Codons per Amino Acid and per Genome in the Code of Man Compared to Other Organisms According to the Rotating Circular Genetic Code. Neuroquantology 9.
- Castro-Chavez F, Vickers KC, Lee JS, Tung CH, Morrisett JD (2013) Effect of Lyso-phosphatidylcholine and schnurri-3 on osteogenic transdifferentiation of vascular smooth muscle cells to calcifying vascular cells in 3D culture. Biochim Biophys Acta 1830 : 3828–3834.
- Vickers KC1, Castro-Chavez F, Morrisett JD (2010) Lyso-phosphatidylcholine induces osteogenic gene expression and phenotype in vascular smooth muscle cells. Atherosclerosis 211: 122-129.
- Castro-Chavez F, Morrisett JD (2011)Osteogenic transdifferentiation of vascular smooth muscle cells to calcifying vascular cells in 3D culture: enhancement by lyso-phosphatidylcholine and attenuation by Schnurri-3. Poster 1108-116, ACC 11, J Am Coll Cardiol 57: E1548.
- Castro-Chavez F, Lee JS, Tung CH, Morrisett JD (2011) Osteogenic transdifferentiation of vascular smooth muscle cells in 3D, detecting hydroxyapatite with a specific probe derived from the alpha region of osteocalcin, Rolanette and Berdon Lawrence Bone Disease Program of Texas Scientific Retreat, Marriott Medical Center Hotel. Baylor College of Medicine and MD Anderson Cancer Center, p3.
- Lee JS1, Morrisett JD, Tung CH (2012) Detection of hydroxyapatite in calcified cardiovascular tissues. Atherosclerosis 224: 340-347.
- Negadi T (2012) the irregular (integer) tetrahedron as a warehouse of biological information. Symm Cult Sci 23(3-4):403-426.
- Rakocevic MM (2013) Harmonic mean as a determinant of the genetic code. arXiv:1305.5103v4 [q-bio.OT], 20 p.
- Negadi T (2011) The multiplet structure of the genetic code, from one and small number. Neuroquantology 9 : 379-384.
- Negadi T (2011) A “quantum-like” approach to the genetic code. Neuroquantology 9 : 785-798.
- Filatov F (2009) A molecular mass gradient is the key parameter of the genetic code organization. arXiv:0907.3537v1 [q-bio.OT], 9 p.
- Mukkamala P, Palvolgyi D (2012) Drawing cubic graphs with the four basic slopes. Lect Notes Comput Sc (7034) 254-265.
- Jeon G, Fang Y, Lee K, Kang K, Jeong J (2012) Improved low-complexity interpolation algorithm for deinterlacing. OJEEE 2 : 259-263.
- Doka G (2002) Life Cycle Assessment of municipal solid waste incineration with the PECK technology ARGE PECK (Zurich) 80 p.
- Bari AT, Reaz MR, Islam AK, Choi HJ, Jeong BS (2013) Effective Encoding for DNA Sequence Visualization Based on Nucleotide's Ring Structure. Evol Bioinform Online 9: 251-261.
- Godina-Nava JJ (2013) Horizontal symmetry in the algebraic approach of genetic code. arXiv:1302.3289v1 [physics.gen-ph], 20 p.
- Mango R, Predazzi IM, Romeo F, Novelli G (2011) LOX-1/LOXIN: the yin/yang of atheroscleorosis. Cardiovasc Drugs Ther 25: 489-494.
- Rissman EF (2008) Roles of oestrogen receptors alpha and beta in behavioural neuroendocrinology: beyond Yin/Yang. J Neuroendocrinol 20: 873-879.
- Weber DS, Griendling KK (2003) The yin/yang of superoxide dismutase mimetics: potential cardiovascular therapies? Br J Pharmacol 139: 1059-1060.
- Mann DL (2002) The yin/yang of innate stress responses in the heart. Cold Spring Harb Symp Quant Biol 67: 363-370.
- Weissmann G, Goldstein I, Hoffstein S, Chauvet G, Robineaux R (1975) Yin/Yang modulation of lysosomal enzyme release from polymorphonuclear leukocytes by cyclic nucleotides. Ann N Y Acad Sci 256: 222-232.
- Clark CE, Liu Y, Cooper HM (2014) The Yin and Yang of Wnt/Ryk axon guidance in development and regeneration. Sci China Life Sci 57: 366-371.
- Wu XS, Wu LG (2014) The yin and yang of calcium effects on synaptic vesicle endocytosis. J Neurosci 34: 2652-2659.
- Martin DS, Grocott MP (2013) III. Oxygen therapy in anaesthesia: the yin and yang of O2. Br J Anaesth 111: 867-871.
- Olsen LF, Issinger OG, Guerra B (2013) The Yin and Yang of redox regulation. Redox Rep 18: 245-252.
- Nie J1, Han X, Shi Y (2013) SAD-A and AMPK kinases: the "yin and yang" regulators of mTORC1 signaling in pancreatic β cells. Cell Cycle 12: 3366-3369.
- Martins-Green M (2013) The Yin and Yang of Integrin Function in Re-Epithelialization During Wound Healing. Adv Wound Care (New Rochelle) 2: 75-80.
- Shiekhattar R (2013) The Yin and Yang of enhancer-like RNAs. EMBO J 32: 2533-2534.
- Han HW, Ohn JH, Moon J, Kim JH (2013) Yin and Yang of disease genes and death genes between reciprocally scale-free biological networks. Nucleic Acids Res 41: 9209-9217.
- Pradere JP, Dapito DH, Schwabe RF (2013) The Yin and Yang of Toll-like receptors in cancer. Oncogene .
- Tobin DM, Ramakrishnan L (2013) TB: the Yin and Yang of lipid mediators. Curr Opin Pharmacol 13: 641-645.
- Stange DE, Clevers H (2013) Concise review: the yin and yang of intestinal (cancer) stem cells and their progenitors. Stem Cells 31: 2287-2295.
- Edgerton DS, Cherrington AD (2013) Glucagon's yin and yang effects on hepatic glucose production. Nat Med 19: 674-675.
- Axelrad MD, Budagov T, Atzmon G (2013) Telomere length and telomerase activity; a Yin and Yang of cell senescence. J Vis Exp : e50246.
- Munro CA (2013) Chitin and glucan, the yin and yang of the fungal cell wall, implications for antifungal drug discovery and therapy. Adv Appl Microbiol 83: 145-172.
- Alvarez AF, Rodriguez C, Georgellis D (2013) Ubiquinone and menaquinone electron carriers represent the yin and yang in the redox regulation of the ArcB sensor kinase. J Bacteriol 195: 3054-3061.
- Tan M, Wang M, Zhou XL, Yan W, Eriani G, et al. (2013) The Yin and Yang of tRNA: proper binding of acceptor end determines the catalytic balance of editing and aminoacylation. Nucleic Acids Res 41: 5513-5523.
- So AY, Zhao JL, Baltimore D (2013) The Yin and Yang of microRNAs: leukemia and immunity. Immunol Rev 253: 129-145.
- Jiménez J, Truman AW, Menoyo S, Kron SJ, Clotet J (2013) The yin and yang of cyclin control by nutrients. Cell Cycle 12: 865-866.
- Zhang EY, Kong KF, Altman A (2013) The yin and yang of protein kinase C-theta (PKCθ): a novel drug target for selective immunosuppression. Adv Pharmacol 66: 267-312.
- Burke AJ, Sullivan FJ, Giles FJ, Glynn SA (2013) The yin and yang of nitric oxide in cancer progression. Carcinogenesis 34: 503-512.
- Vij N, Downey GP (2013) The yin and yang of cystic fibrosis transmembrane conductance regulator function: implications for chronic lung disease. Am J Respir Crit Care Med 187: 120-122.
- Vasquez KM, Wang G (2013) The yin and yang of repair mechanisms in DNA structure-induced genetic instability. Mutat Res 743-744: 118-31.
- Kubli DA, Gustafsson ĹB (2012) Mitochondria and mitophagy: the yin and yang of cell death control. Circ Res 111: 1208-1221.
- Weinberg A1, Jin G, Sieg S, McCormick TS (2012) The yin and yang of human Beta-defensins in health and disease. Front Immunol 3: 294.
- Hein GJ, Baker C, Hsieh J, Farr S, Adeli K (2013) GLP-1 and GLP-2 as yin and yang of intestinal lipoprotein production: evidence for predominance of GLP-2-stimulated postprandial lipemia in normal and insulin-resistant states. Diabetes 62 : 373-381.
- Adams EJ, Luoma AM (2012) The yin and yang of CD1d recognition. Nat Immunol 13: 814-815.
- Sladek FM (2012) The yin and yang of proliferation and differentiation: cyclin D1 inhibits differentiation factors ChREBP and HNF4α. Cell Cycle 11: 3156-3157.
- Hou J (2012) The yin and yang of claudin-14 function in human diseases. Ann N Y Acad Sci 1258: 185-190.
|
Tables at a glance
 |
| Table 1 |
Figures at a glance
 | |
 | |
 | |
 | |
 | |
 |
| Figure 1 | |
Figure 2 | |
Figure 3 | |
Figure 4 | |
Figure 5 | |
Figure 6 |
 | |
 | |
 | |
 | |
 | |
 |
| Figure 7 | |
Figure 8 | |
Figure 9 | |
Figure 10 | |
Figure 11 | |
Figure 12 |
 | |
 | |
 | |
 | |
 | |
 |
| Figure 13 | |
Figure 14 | |
Figure 15 | |
Figure 16 | |
Figure 17 | |
Figure 18 |
 | |
 | |
 | |
 | |
 | |
 |
| Figure 19 | |
Figure 20 | |
Figure 21 | |
Figure 22 | |
Figure 23 | |
Figure 24 |
 | |
 | |
 |
| Figure 25 | |
Figure 26 | |
Figure 27 |
|
|
|